In 1922, 14-year-old Leonard Thompson became the first patient to receive insulin, marking a defining milestone in the emergence of peptide therapeutics. Today, as peptide molecules become increasingly complex, they have evolved from simple natural sequence mimics into highly engineered structures with precise chemical modifications and well-defined three-dimensional structures.
Yet as molecular complexity increases, technologies such as solid phase synthesis, phage and mRNA display remain limited in their ability to fully explore chemical space.
To address these challenges and support the growing global demand for peptide drug discovery, BioDuro has built an advanced Peptide DNA-Encoded Libraries (DEL) platform designed to advance more efficient peptide discovery. We recently spoke with our Director of Peptide DEL Platform, Dr. Yihui Xie, about the platform and how it advances peptide discovery.
The industry is moving beyond volume-driven metrics. While phage and mRNA display achieve large library sizes, they rely on biological translation systems and face key constraints:
Limitations in Chemical Space: These libraries are restricted to the 20 natural amino acids and typically allow only 1-5 non-canonical amino acids (ncAAs) per peptide. In modern drug discovery, this level of diversity is often not sufficient to achieve sufficient molecular differentiation to support viable candidate development
Challenges in Lead Optimizations and Oral Bioavailability: Because these libraries consist of natural linear peptides predominantly, hits often show low oral bioavailability. As a result, researchers frequently need to introduce intensive synthetic modifications after screening, which can reduce binding affinity and lead to challenges during lead optimization.
Limited structural control: Within a biological system, researchers have limited ability to pre-program more complex architectures, such as site-specific macrocyclization or constrained scaffolds due to the lack of orthogonal chemistry.
DEL technology addresses these limitations through a fully chemical synthesis approach, enabling broad incorporation of ncAAs, including D-amino acids and N-methyl amino acids, at defined positions. At BioDuro, we prioritize a curated library exceeding 10^9 compounds, focusing on quality over scale. This approach ensures hits are designed with strong optimization potential and drug-like properties from the outset.
Our platform integrates high-quality oligopeptide building blocks, diverse chemistries, and rational library design to deliver more reliable and structurally complex peptide discovery.
We apply strict analytical validation for our library of over 1,000 unnatural amino acid and oligopeptide building blocks, ensuring purity levels consistently exceed 90%. Coupled with HPLC purification across the library synthesis, this chemical precision results in more reliable hit identification, as it significantly reduces the risk of failure during later lead optimization stages.
In terms of design flexibility, we offer a peptide length range of 4 to 20 amino acids. This broad span allows us to customize libraries for specific target needs, whether the goal is a small peptide with high permeability or a large molecule designed to disrupt complex protein-protein interactions (PPIs). By combining this range with over 10 cyclization strategies, we can build stable constrained scaffolds. These rigid structures are essential for enhancing metabolic stability and improving the oral bioavailability of peptide drugs.
Finally, the platform's strength lies in the integration of Peptide DEL with our Direct-to-Biology (D2B) validation system. High-potential hits from DEL screening move directly into parallel synthesis at scale, enabling immediate functional testing without purification or sample handling. This screen-to-test workflow improves turnaround time by up to 52-fold compared to traditional synthesis, while reducing cost by over 70%+ per peptide.
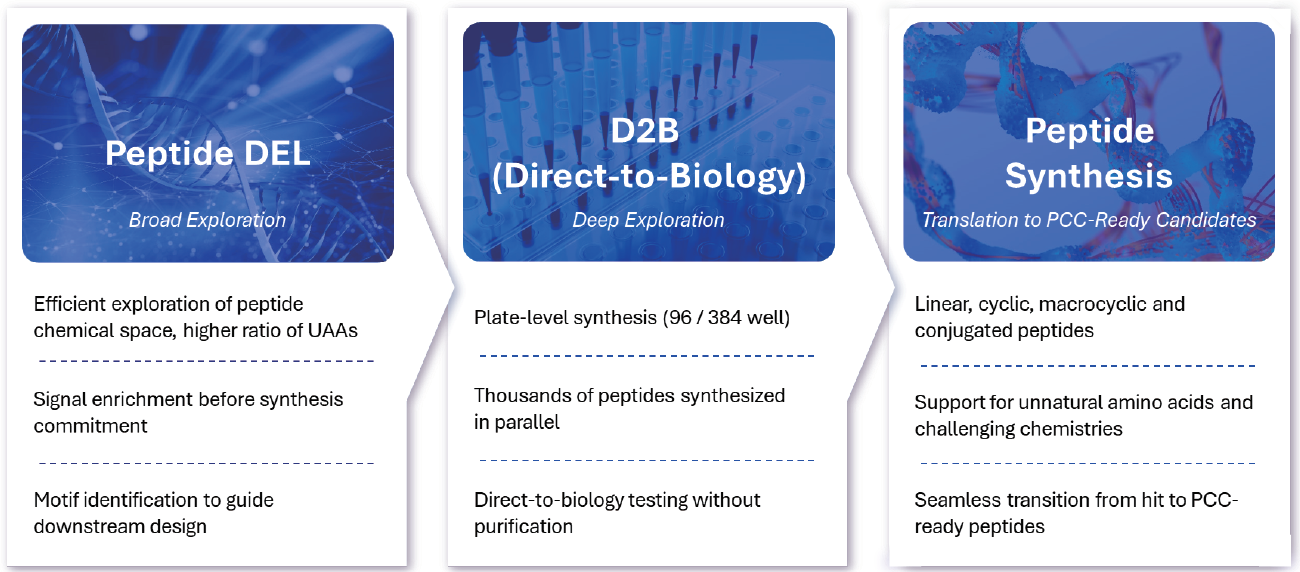
A seamless workflow transforming Peptide DEL hits into PCC-ready candidates
With an 80–90% synthesis success rate for both linear and cyclic peptides, we can improve potency from initial μM hits to ~30 nM within 8–12 weeks, supporting a clear path toward preclinical discovery.
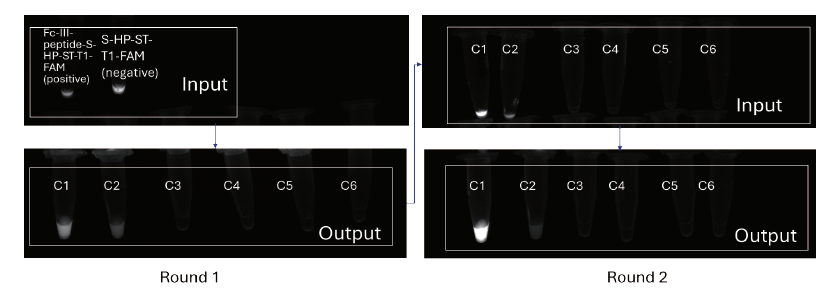
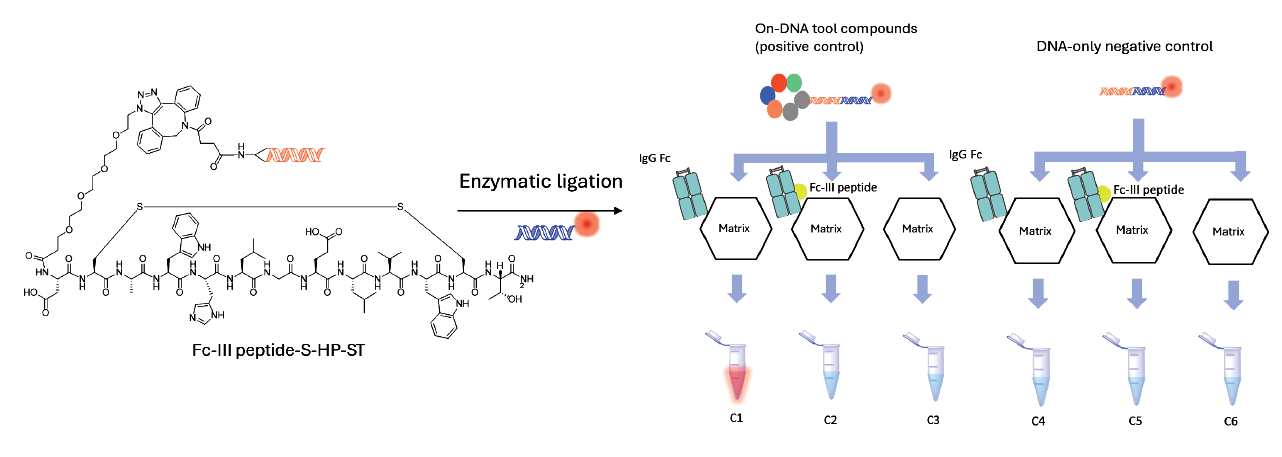

Fluorescent images of Eluates
In a recent validation study targeting the IgG Fc protein, we showed the platform's ability to identify target sequences from a library of thousands of compounds. Fluorescence experiment showed a clear enrichment of target binders: after two rounds of selection, the known positive peptide sequences increased significantly. This enrichment highlights the platform's ability to isolate true signals from a large pool of candidates with high efficiency.
The case study demonstrated strong screening purity. In control experiments, there was minimal interference from matrix or DNA background signals. This low-noise performance is important in early-stage discovery, as it increases confidence in identified hits and reduces the risk of false positives during follow-up validation. Overall, our platform provides reliable starting points for downstream discovery.
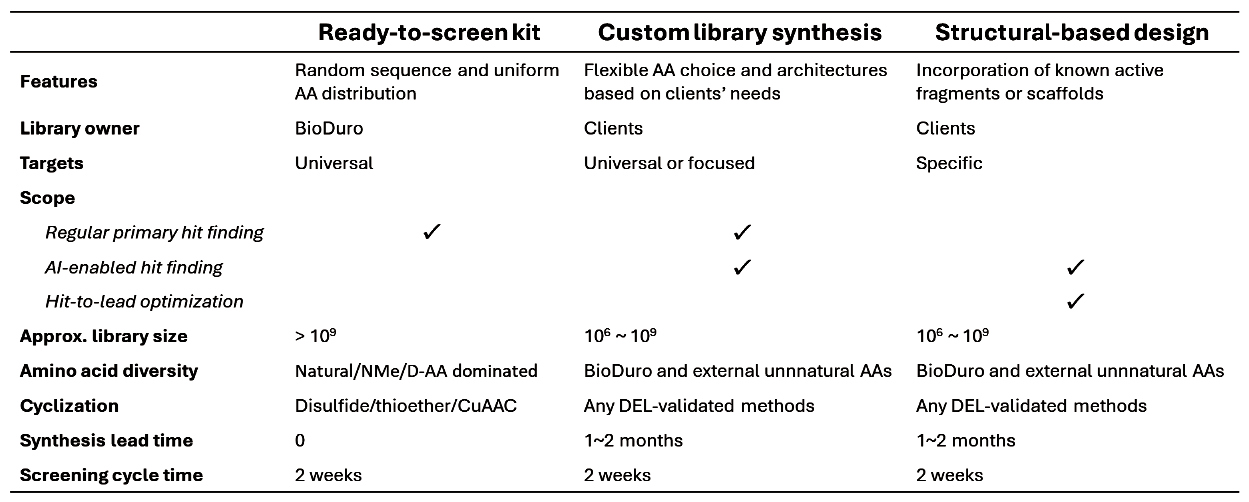
Tailored to Your Discovery Goals
Whether starting with a target for rapid hit identification or optimizing an existing scaffold, our three engagement models support programs across different stages of peptide discovery.